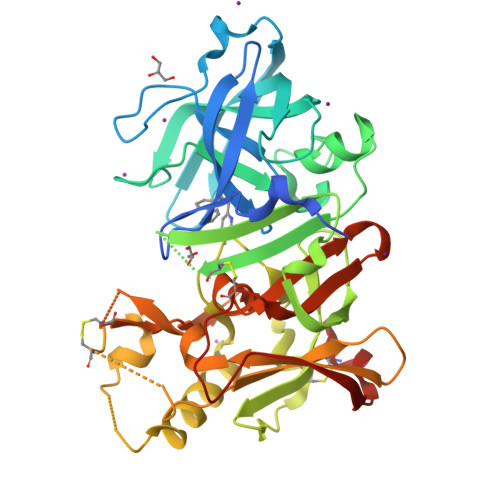
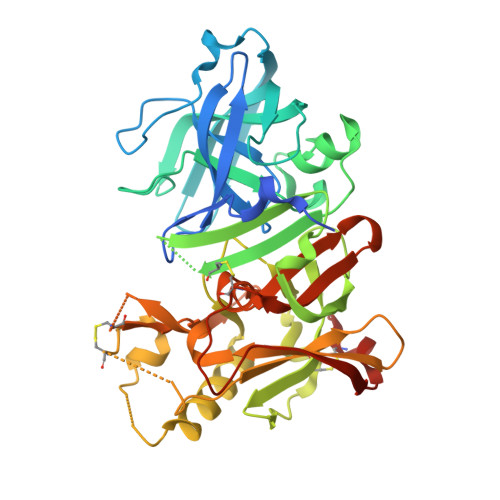
Conformational restriction approach to beta-secretase (BACE1) inhibitors: effect of a cyclopropane ring to induce an alternative binding mode.
Yonezawa, S., Yamamoto, T., Yamakawa, H., Muto, C., Hosono, M., Hattori, K., Higashino, K., Yutsudo, T., Iwamoto, H., Kondo, Y., Sakagami, M., Togame, H., Tanaka, Y., Nakano, T., Takemoto, H., Arisawa, M., Shuto, S.(2012) J Med Chem 55: 8838-8858
- PubMed: 22998419
- DOI: https://doi.org/10.1021/jm3011405
- Primary Citation of Related Structures:
3VV6, 3VV7, 3VV8 - PubMed Abstract:
Improvement of a drug's binding activity using the conformational restriction approach with sp³ hybridized carbon is becoming a key strategy in drug discovery. We applied this approach to BACE1 inhibitors and designed four stereoisomeric cyclopropane compounds in which the ethylene linker of a known amidine-type inhibitor 2 was replaced with chiral cyclopropane rings. The synthesis and biologic evaluation of these compounds revealed that the cis-(1S,2R) isomer 6 exhibited the most potent BACE1 inhibitory activity among them. X-ray structure analysis of the complex of 6 and BACE1 revealed that its unique binding mode is due to the apparent CH-π interaction between the rigid cyclopropane ring and the Tyr71 side chain. A derivatization study using 6 as a lead molecule led to the development of highly potent inhibitors in which the structure-activity relationship as well as the binding mode of the compounds clearly differ from those of known amidine-type inhibitors.
Organizational Affiliation:
Shionogi Innovation Center for Drug Discovery, Shionogi & Co., Ltd., Kita-21 Nishi-11 Kita-ku, Sapporo 001-0021, Japan. shu@pharm.hokudai.ac.jp