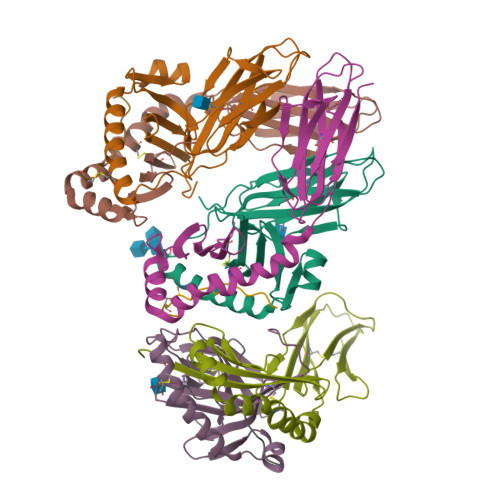
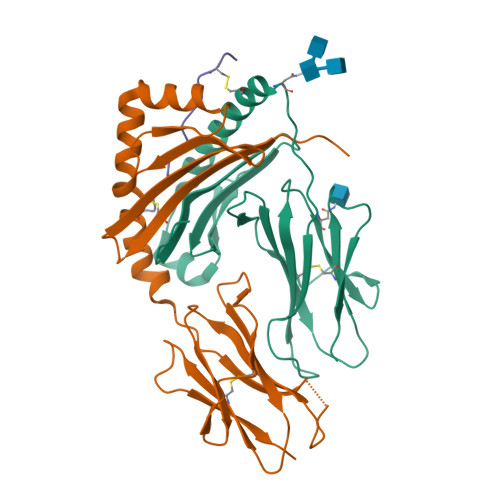
C-terminal modification of the insulin B:11-23 peptide creates superagonists in mouse and human type 1 diabetes.
Wang, Y., Sosinowski, T., Novikov, A., Crawford, F., Neau, D.B., Yang, J., Kwok, W.W., Marrack, P., Kappler, J.W., Dai, S.(2018) Proc Natl Acad Sci U S A 115: 162-167
- PubMed: 29255035
- DOI: https://doi.org/10.1073/pnas.1716527115
- Primary Citation of Related Structures:
5UJT, 6BLQ, 6BLR, 6BLX - PubMed Abstract:
A polymorphism at β57 in some major histocompatibility complex class II (MHCII) alleles of rodents and humans is associated with a high risk for developing type 1 diabetes (T1D). However, a highly diabetogenic insulin B chain epitope within the B:9-23 peptide is presented poorly by these alleles to a variety of mouse and human CD4 T cells isolated from either nonobese diabetic (NOD) mice or humans with T1D. We have shown for both species that mutations at the C-terminal end of this epitope dramatically improve presentation to these T cells. Here we present the crystal structures of these mutated peptides bound to mouse IA g7 and human HLA-DQ8 that show how the mutations function to improve T-cell activation. In both peptide binding grooves, the mutation of B:22R to E in the peptide changes a highly unfavorable side chain for the p9 pocket to an optimal one that is dependent on the β57 polymorphism, accounting for why these peptides bind much better to these MHCIIs. Furthermore, a second mutation of the adjacent B:21 (E to G) removes a side chain from the surface of the complex that is highly unfavorable for a subset of NOD mouse CD4 cells, thereby greatly enhancing their response to the complex. These results point out the similarities between the mouse and human responses to this B chain epitope in T1D and suggest there may be common posttranslational modifications at the C terminus of the peptide in vivo to create the pathogenic epitopes in both species.
Organizational Affiliation:
Department of Biomedical Research, National Jewish Health, Denver, CO 80206.