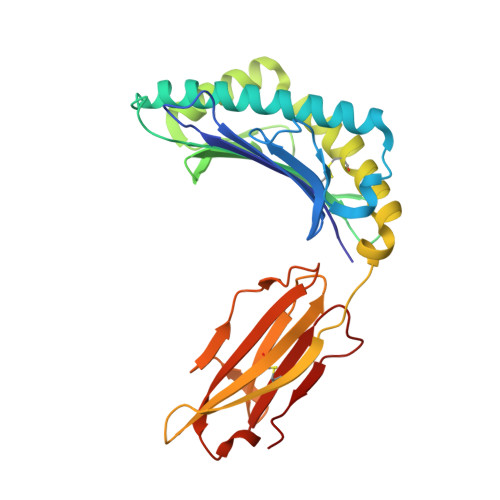
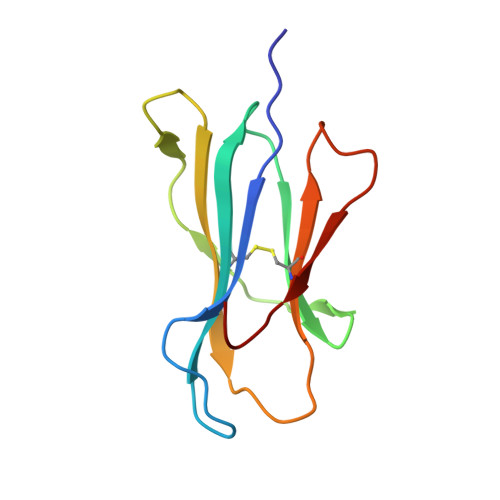
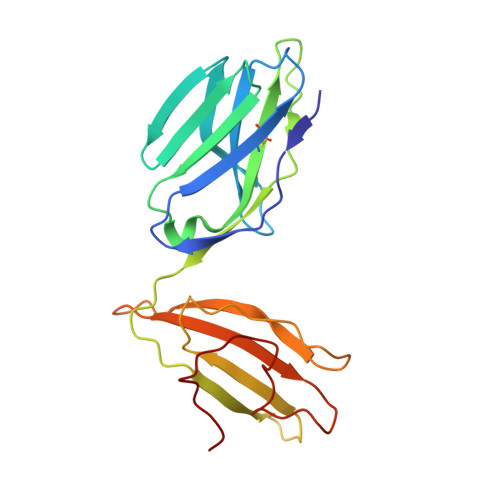
Hotspot autoimmune T cell receptor binding underlies pathogen and insulin peptide cross-reactivity.
Cole, D.K., Bulek, A.M., Dolton, G., Schauenberg, A.J., Szomolay, B., Rittase, W., Trimby, A., Jothikumar, P., Fuller, A., Skowera, A., Rossjohn, J., Zhu, C., Miles, J.J., Peakman, M., Wooldridge, L., Rizkallah, P.J., Sewell, A.K.(2016) J Clin Invest 126: 2191-2204
- PubMed: 27183389
- DOI: https://doi.org/10.1172/JCI85679
- Primary Citation of Related Structures:
5C07, 5C08, 5C09, 5C0A, 5C0B, 5C0C, 5C0D, 5C0E, 5C0F, 5C0G, 5C0I, 5C0J, 5N1Y - PubMed Abstract:
The cross-reactivity of T cells with pathogen- and self-derived peptides has been implicated as a pathway involved in the development of autoimmunity. However, the mechanisms that allow the clonal T cell antigen receptor (TCR) to functionally engage multiple peptide-major histocompatibility complexes (pMHC) are unclear. Here, we studied multiligand discrimination by a human, preproinsulin reactive, MHC class-I-restricted CD8+ T cell clone (1E6) that can recognize over 1 million different peptides. We generated high-resolution structures of the 1E6 TCR bound to 7 altered peptide ligands, including a pathogen-derived peptide that was an order of magnitude more potent than the natural self-peptide. Evaluation of these structures demonstrated that binding was stabilized through a conserved lock-and-key-like minimal binding footprint that enables 1E6 TCR to tolerate vast numbers of substitutions outside of this so-called hotspot. Highly potent antigens of the 1E6 TCR engaged with a strong antipathogen-like binding affinity; this engagement was governed though an energetic switch from an enthalpically to entropically driven interaction compared with the natural autoimmune ligand. Together, these data highlight how T cell cross-reactivity with pathogen-derived antigens might break self-tolerance to induce autoimmune disease.